Last year, we were working with a mid-sized clinical genetics lab on a bioinformatics consulting project. During one of our discussions, their lead geneticist mentioned, "We have a three-week backlog on variant interpretation.” This wasn't a resource problem they could solve by hiring more people. The issue was deeper. Their small bioinformatics team was spending entire days running variants through different annotation tools, cross-referencing and then manually checking gnomAD frequencies, then looking up each significant variant in OMIM and PubMed. The whole process was taking 2-4 weeks per sample, and the bottleneck was growing.
At disease-targeted testing laboratories, even with semi-automated tools, variant assessment averages 40 minutes per variant, 22 minutes when literature review is excluded. This labor translates directly into cost. While sequencing itself has approached the "$1,000 genome" threshold, the infrastructure costs for variant interpretation remain substantial, and interpretation costs are unlikely to fall as rapidly as raw sequencing costs.
We spent several weeks talking with clinical genetics teams, observing their workflows firsthand. The inefficiencies weren't surprising, but the scale was.
Workflow studies revealed, genetic counselors report spending only 20–25% of their time in direct face-to-face patient care. The most time goes to patient-related activities, case preparation, follow-up, administrative tasks, and rest goes to activities unrelated to direct patient care.
One genetic counselor we spoke with put it plainly: "I became a genetic counselor to help families understand their risks and make informed decisions. Instead, I spend most of my time chasing down bioinformatics teams and trying to interpret spreadsheets."
The real cost isn't just measured in labor hours. It's measured in delayed diagnoses, missed treatment opportunities, and patients waiting while their data sits in analysis queues.
We set out to answer a straightforward question: what if interpreting genomic data required uploading a file and clicking "Analyze"? The platform we developed handles the complete tertiary analysis workflow:
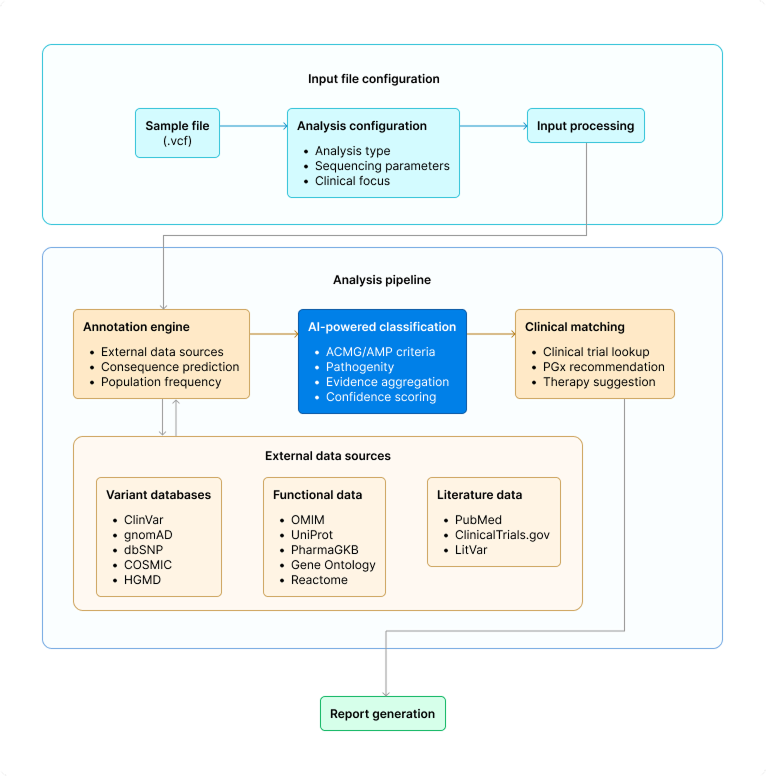
Tertiary Analysis Platform Architecture
Genomic medicine should be accessible to any clinic that wants to use it, not just major academic medical centers with specialized bioinformatics teams.
A small community hospital should be able to offer genomic testing to their cancer patients. A solo genetic counseling practice should be able to provide carrier screening without outsourcing to expensive lab services with long turnaround times. The technology for sequencing is already there, affordable and widespread. The bottleneck is interpretation. Removing that bottleneck makes genomic medicine practical for many more patients who could benefit from it.
The platform roadmap includes:
Rare disease patients often wait years for diagnosis. Phenotype-driven variant prioritization can significantly reduce the time to identify causal variants.
Comparing proband, mother, and father genomes enables detection of de novo mutations that would be impossible to identify from singleton analysis.
Inversions, translocations, and large copy number variants require different analytical approaches than SNVs and small indels.
For common diseases where risk is distributed across many variants of small effect, aggregate scoring provides clinically relevant risk stratification.
Seamless clinical workflows require genomic results to flow directly into electronic medical records without manual transcription.
Multidisciplinary tumor boards and rare disease case conferences benefit from shared access to variant evidence and classification rationale.
Alternatively, labs build custom systems and become dependent on a small internal engineering team. When key developers leave, the platform becomes unmaintainable.
References